Harvard researchers have created predictive software that can precisely identify the most effective ways to target genes with the gene editing mechanism CRISPR-Cas9
(BOSTON) — The remarkable ease and accuracy with which scientists can alter genomes using the CRISPR-Cas9 system has led to promising advances towards improving human health and the environment through genetic engineering. Cas9, a protein found naturally in certain bacteria, functions like a pair of molecular scissors to precisely cut sections of DNA and is extremely effective as a gene editing tool. It can be directed to a specific gene through the use of a matching guide RNA sequence to perform gene mutations, putting programmable control of gene editing in the hands of scientists.
But even though Cas9 can precisely carry out these functions, more than one guide RNA can be a possible match for any given gene target, which leaves scientists with preliminary troubleshooting work to select the best guide RNA for any given gene editing task.
To eliminate the trial-and-error process of selecting guide RNAs for each job, a team led by Harvard geneticist George Church, Ph.D., who is a Core Faculty member at the Wyss Institute for Biologically Inspired Engineering, the Robert Winthrop Professor of Genetics at Harvard Medical School, and Professor of Health Sciences and Technology at Harvard and MIT, has developed a new, straightforward software program for predicting the best guide RNAs to direct Cas9 to gene targets. The work is reported in the July 13 issue of Nature Methods.
“We started off by asking ourselves, is there something in the guide RNA sequence that could suggest one would work better than others?” said the study’s lead author Raj Chari, Ph.D., Postdoctoral Fellow in Genetics at Harvard Medical School.
The software hierarchically ranks how effective any given guide RNA will target a desired gene target based on experimental data gathered using human genomes, unlike other gene targeting algorithms currently available to scientists that operate on first principles only.
“This will allow for widespread improvement in the speed and accuracy with which scientists can select the appropriate guide RNA for achieving their desired gene editing results,” said Church.
Made publicly available by Church’s team, it not only contains information about effective guide RNAs for Cas9 extracted from the bacteria species S. pyogenes, which is the most commonly-used bacterial source of Cas9, but also includes the world’s most comprehensive library of guide RNA from a different bacteria species that has been rising in its popularity of use, S. thermophilus.
To develop the algorithm that is the basis for the new software, Church’s team carried out a high-throughput analysis of the activity between many gene targets and complementary guide RNAs, searching for patterns in the guide RNA sequences that could indicate how effectively they would bind to any given gene target.
In doing so, they chose a unique sequence from each gene target and synthesized an identical strand of DNA, compiling the DNA into a large library representing all of the targets. The gene targets were then inserted into the genomes of cultured human cells. To test which guide RNAs would be the most effective at reaching each of these unique targets, complementary guide RNAs were inserted with Cas9 into the cells, giving Cas9 the opportunity to make genome mutations at the sites where successful matching occurred. A basic genome extraction and sequencing process then easily revealed which guide RNA was the best match for each target.
From the data the team developed their novel algorithm to rank and score the most effective guide RNAs for targeting virtually any human gene, even ones lacking experimental data in their software database.
“Analyzing our results, we identified certain features in guide RNA sequences that are indicative of how well they work at directing Cas9 to a desired gene target,” said Chari. “By designing an algorithm based on these features, we can now look at any guide RNA sequence and assign it a score for how effective it is predicted to work.”
By speeding up this part of the gene editing process, scientists can focus efforts on applying Cas9 gene editing to the development of not only gene therapies against diseases such as sickle cell anemia, cancer, or HIV.
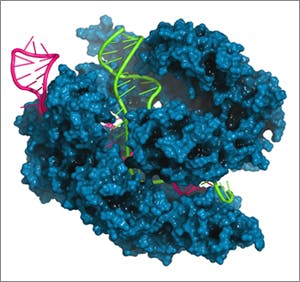
“For any scientist using RNA-guided Cas9 in their day to day work, it will be greatly beneficial to have a tool like this that quickly narrows down which guide RNA will be the most effective to direct Cas9 to a gene of interest,” said Chari.
“New tools like this are making gene editing even more effortless and programmable,” said Wyss Institute Founding Director Donald Ingber, M.D., Ph.D., who is also the Judah Folkman Professor of Vascular Biology at Harvard Medical School and Boston Children’s Hospital and Professor of Bioengineering at Harvard SEAS. “Being able to quickly select the best guide RNA will accelerate efforts in a broad range of areas, including identification of novel gene targets for development of new therapeutics.”